Research Focus
I am focused on developing predictive models of biological systems, in the intersection of machine learning, synthetic biology and automation. My interests range from synthetic biology, machine learning, systems biology, metabolic flux analysis, data visualization, scientific software development, ecology, and automation and complexity.
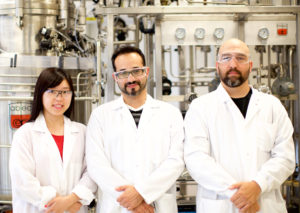
In my current position at the Joint BioEnergy Institute, I develop predictive quantitative models of microbial metabolism to direct metabolic engineering efforts, and use them to (e.g.) improve biofuel yields or design microbiomes.
Our efforts are divided among machine learning, flux modeling approaches, retrobiosynthesis, automation and software development for visualization and acquisition of data. In the following links, you can find more detailed information on my research interests and the full story of how a physicist ended up working in biology.
Projects
- Machine learning and data mining for pathway engineering.
- Metabolic flux analysis for host engineering.
- Retrobiosynthesis.
- Multiomics data visualization, capture and storage.
Featured Media
Could Synthetic Biology Stop Global Warming? (NPR)
What Termites Can Teach Us (New Yorker)
Underbug (chapter 12)
New Machine Learning Approach Could Accelerate Bioengineering
Hector Garcia interviewed by California-Spain Chamber of Commerce
Navigating an Ocean of Biological Data in the Modern Era
JBEI Expert Hector Garcia on how metabolic engineering is applied to biofuel production
Unlocking Clean Cheap Energy
¿Para cuándo los biocarburantes de 2ª generación?
Multicultural Metabolic Map
Leaving Out the Details
Featured Publications
- “Common principles and best practices for engineering microbiomes.”, Nature Reviews Microbiology (2019)
- “An automated ‘cells-to-peptides’ sample preparation workflow for high-throughput, quantitative proteomic assays of microbes.”, Journal of proteome research (2019)
- “Opportunities at the Intersection of Synthetic Biology, Machine Learning, and Automation.”, ACS Synthetic Biology (2019)
- “Lessons from two Design-Build-Test-Learn cycles of dodecanol production in Escherichia coli aided by machine learning.”, ACS synthetic biology (2019)
- “Parallel Integration and Chromosomal Expansion of Metabolic Pathways.”, ACS Synthetic Biology (2018).
- “A machine learning approach to predict metabolic pathway dynamics from time-series multiomics data.”, Nature pj Systems Biology & Applications (2018).
- “Automated flow-based/digital microfluidic platform integrated with onsite electroporation process for multiplex genetic engineering applications.”, Micro Electro Mechanical Systems (MEMS): IEEE (2018).
- “Industrial brewing yeast engineered for the production of primary flavor determinants in hopped beer.” Nature Communications (2018).
- “Constraining Genome-Scale Models to Represent the Bow Tie Structure of Metabolism for 13C Metabolic Flux Analysis.”, Metabolites (2018).
- “ClusterCAD: a computational platform for type I modular polyketide synthase design”, Nucleic Acid Research (2017)
- “The Experiment Data Depot: a web-based software tool for biological experimental data storage, sharing, and visualization”, ACS Synthetic Biology (2017)
- “The JBEI quantitative metabolic modeling library (jQMM): a python library for modeling microbial metabolism”, BMC Bioinformatics (2017)
- “13C Metabolic Flux Analysis for Systematic Metabolic Engineering of S. cerevisiae for Overproduction of Fatty Acids”, Frontiers in Bioengineering Biotechnology (2016)
- “Synthetic and systems biology for microbial production of commodity chemicals”, Nature pj Systems Biology & Applications (2016)
- “A Method to Constrain Genome-Scale Models with 13C Labelling Data”, PLoS Computational Biology (2015)
- “Principal component analysis of proteomics (PCAP) as a tool to direct metabolic engineering”, Metabolic Engineering (2015)
- “A peptide-based method for 13C Metabolic Flux Analysis in microbial communities”, PLoS Computational Biology (2014)
- “Advances in analysis of microbial metabolic fluxes via (13)C isotopic labeling”, Mass Spectrometry Reviews (2009)
- “Integrating ecology into biotechnology”, Current Opinion in Biotechnology (2007)
- “Dissecting biological “dark matter” with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth”, P.N.A.S (2007)
- “Accurate phylogenetic classification of variable-length DNA fragments”, Nature Methods (2007)
- “Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite”, Nature (2007)
- “On the origin and robustness of power-law species-area relationships in ecology”, P.N.A.S (2006)
- “Metagenomic analysis of two enhanced biological phosphorus removal (EBPR) sludge communities”, Nature Biotechnology (2006)