–By Irina Silva
Development of robust microbial platforms for bioproduction requires strains that have been engineered to have efficient carbon uptake, energy generation, tolerance to biomass pretreatment byproducts and the export of final product. Many of these optimizations require expression and overexpression of native and heterologous membrane proteins. Over the years, scientists at the U.S. Department of Energy’s Joint BioEnergy Institute (JBEI), and others have successfully found many such engineering targets. However, JBEI has also found that expression of membrane proteins is challenging and can also impact the microbial growth, thus negatively limiting the use of these discoveries.
In a new paper published in Nature Scientific Reports, Biofuels and Bioproducts Division scientists at JBEI, hypothesized that they could find loci in the genome that could be edited to enhance expression of any given membrane protein of interest.
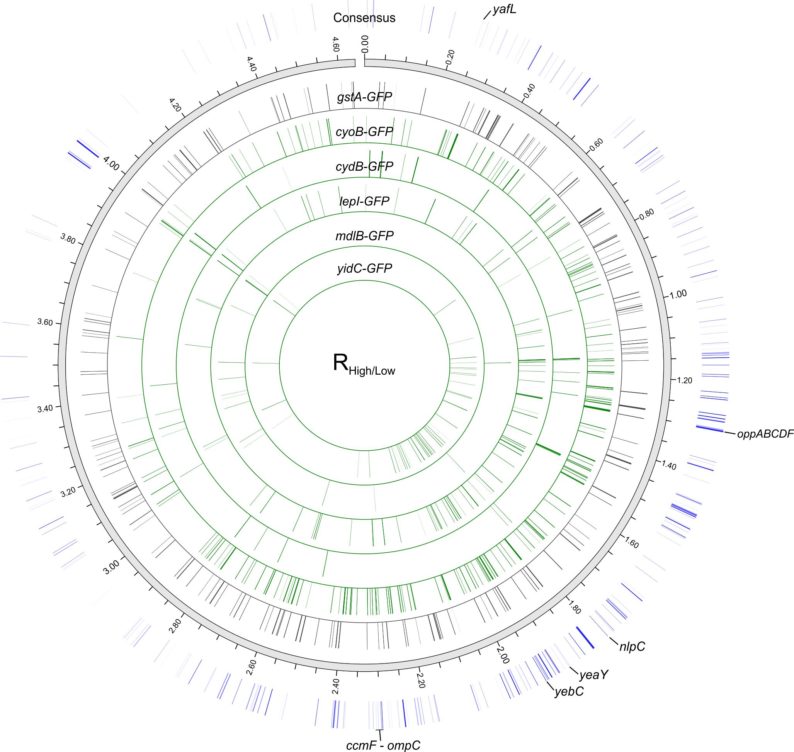
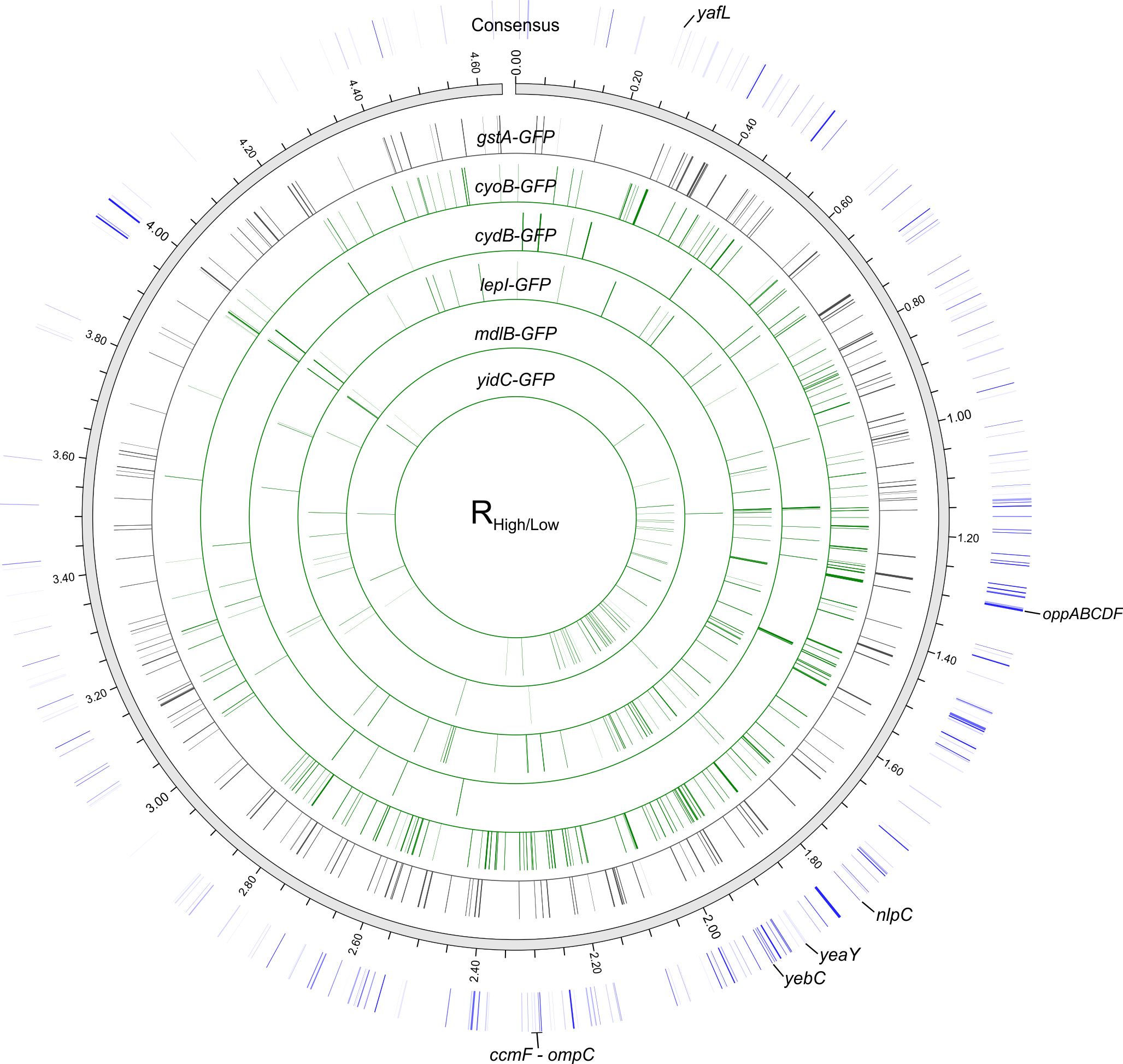
The hypothesis was tested by using a bar coded gene knockout library in E. coli in conjunction with expression of fluorescent protein tagged membrane proteins. Numerous loci in the genome were identified that when disrupted allowed improved expression, and consequently function, of the proteins of interest. These are host backgrounds that are better suited to serve as the conversion platforms. This study also represents a systematic evaluation of a microbial host genome to improve membrane protein expression.
Published in Nature Scientific Reports, “Improving membrane protein expression and function using genomic edits” is authored by Heather M. Jensen, Thomas Eng, Victor Chubukov, Robin A. Herbert and Aindrila Mukhopadhyay.